
|
Help
|
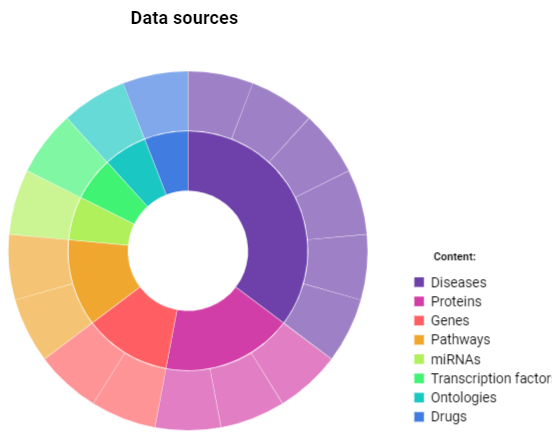
GeDiPNeT is a comprehensive database on all human gene disease pathway networks. The homepage provides a short description of the content of the database and it’s algorithms. This database can be used to retrieve information on human genes and its annotations, SNPs, diseases and pathways. It has been interlinked with various other major databases. Algorithms available in GeDiPNet could be used for performing high-throughput data analysis that could lead to novel insights and hypothesis generation for gene-disease-pathway associations. Data can be retrieved using the navigation tabs (menu bar) on the top of the webpage. Infographic charts as an image are also present on the homepage of GeDiPNeT representing data available in the database. Detailed dynamics charts can be accessed and downloaded using the hyperlinked page on the image. The above figure shows that GeDiPNeT is made up of 17 data sources, has 7201 data on diseases, 12418 gene-disease associations and 66804 literature evidences.
The above chart represents the data sources of GeDiPNeT with respect to each content. The inner circle represents each content as a part of the whole, while the outer circle represents the divisions of each fraction of a particular content. The above histogram represents the distribution of SNPs on chromosomes. The X-axis represents the chromosome number and Y-axis represents the number of SNPs in each chromosome. The chromosomes are arranged in the order of one having the highest number of SNPs. A panel of 6 interactive lists representing data on genes and associated diseases, pathways and associated diseases, diseases and associated genes, transcription factors and target genes and, miRNAs and the target genes;is also available on the homepage of GeDiPNet. 2. Search There are two types of search options in GeDiPNeT.
To build a query, one field (explained below) at a time should be selected and a keyword corresponding to the same should be typed in the blank space. Click on the
Note: Queries can be built combining options 'AND', 'OR' from one section at a time
3. Browse 
Browse enables users to surf the database for genes, pathways and diseases. The information available on each page is explained below:
All the above-mentioned pages can be filtered, queried using keywords, and browsed by alphabets. One or more of these can be clubbed to get more specific results. There are four different analysis that can be performed using GeDiPNet. 
4.1 Enrichment Analysis
Using this tool, user can enter or upload a gene list, which can be used to perform pathway enrichment analyses, disease enrichment analyses (using genes & using functions) The gene list can be a comma separated or new line separated. Pathway enrichment or disease enrichment can be performed using the respective All correct and incorrect entries will be displayed as accepted and rejected gene symbols on the webpage as soon as the gene set is submitted for analysis. 
One can either proceed or go back as per their query. On proceeding, users are directed to either a pathway enrichment page or a disease enrichment page.
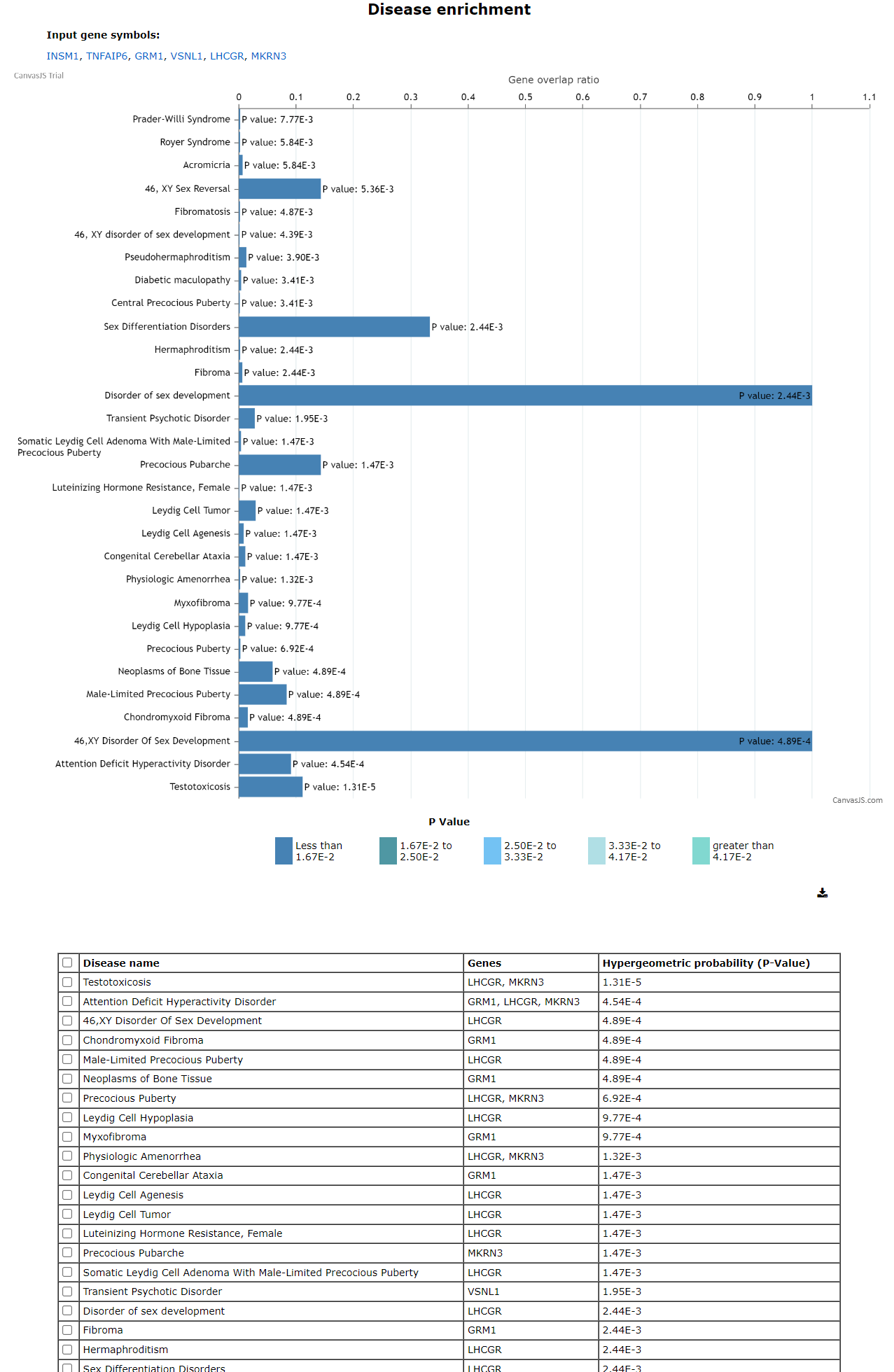
The pathway enrichment page displays the input gene symbols followed by two buttons i.e. The graph is plotted for gene overlap ratio vs disease name for each disease, while showing the p-value for each disease.The hypergeometric probabilityand overlap ratio scores are provided in the table. 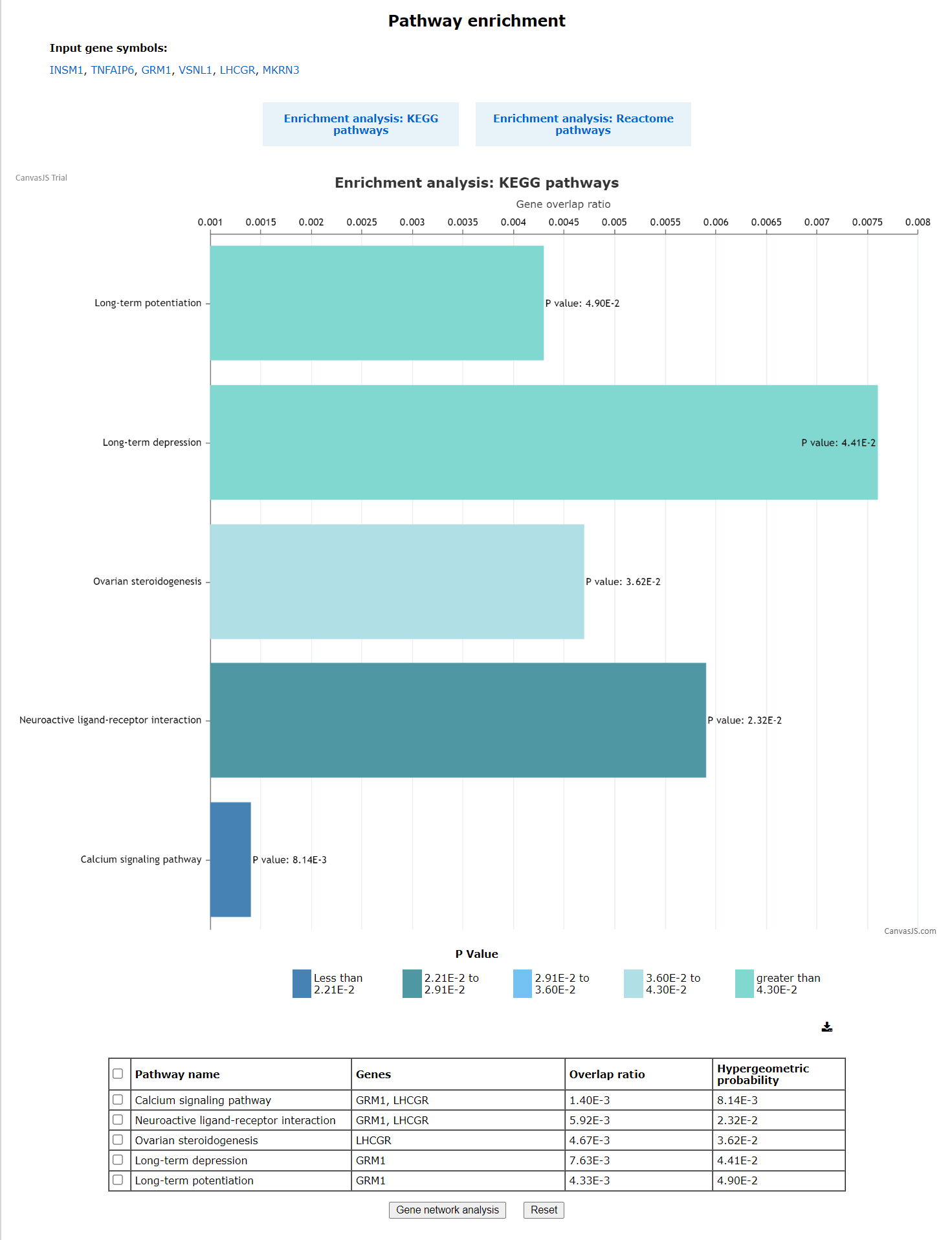
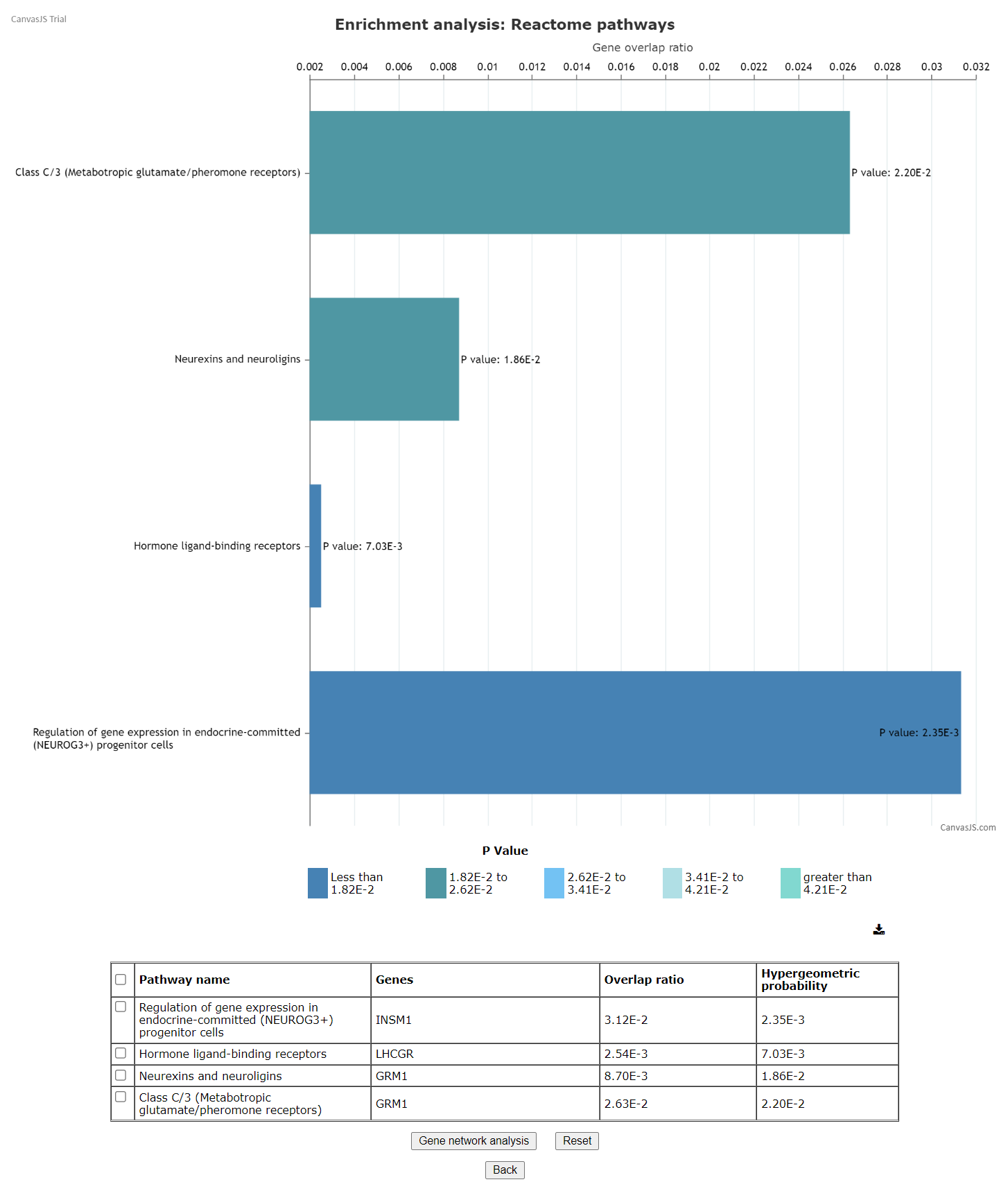
These results can be further taken to gene network analysis using the 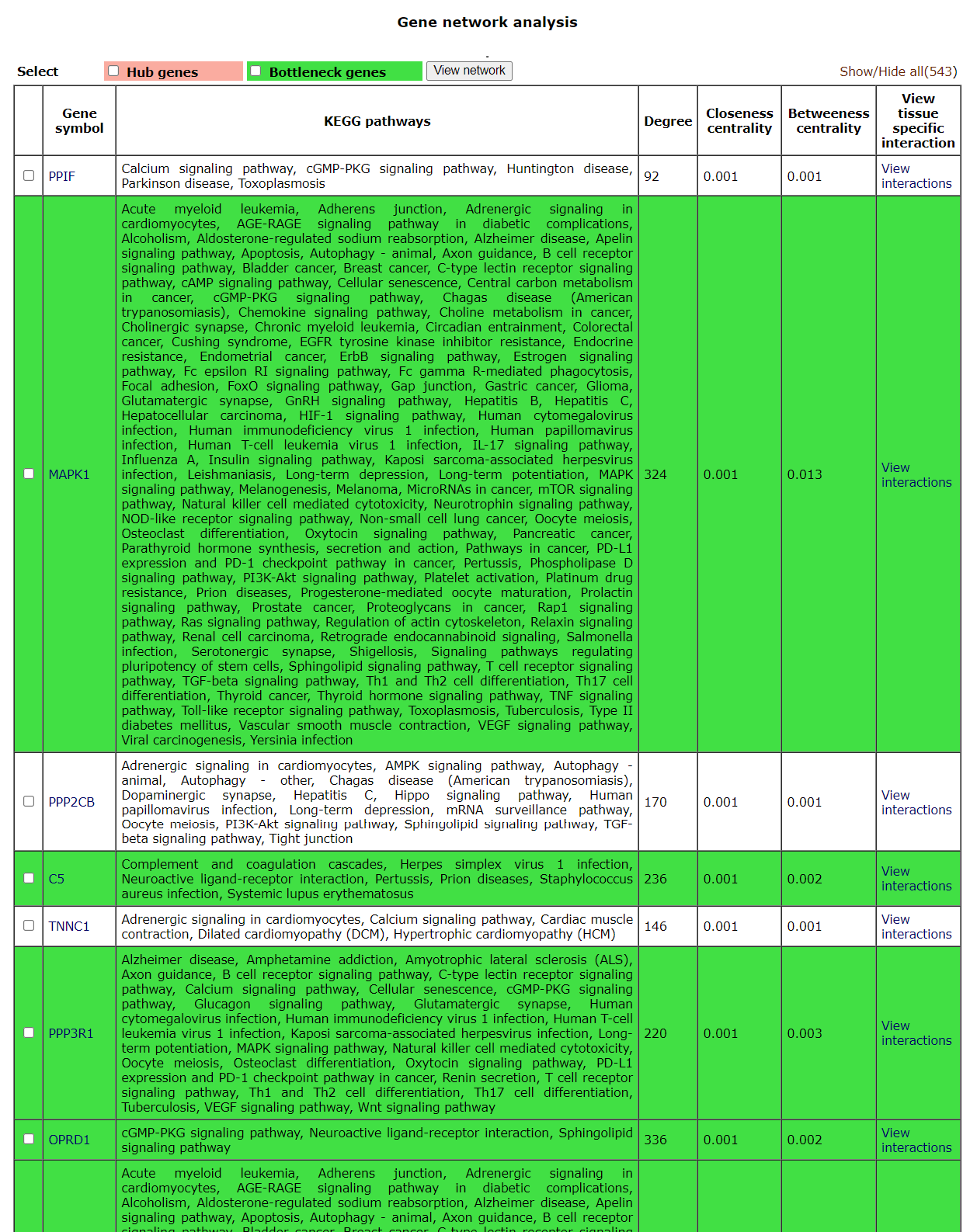
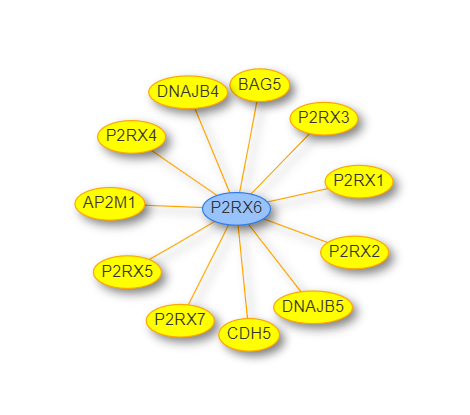
The Users can select the genes in the list and view a network by using the
A "view interaction" button under the "View tissue specific interaction" column in the table. By clicking on it, a tissue specific interaction network is obtained.
Disease enrichment analysis has a similar input and result page as pathway enrichment analysis. One can enter the genelist in the input page and click on the disease enrichment button.
The results can be further analysed using the
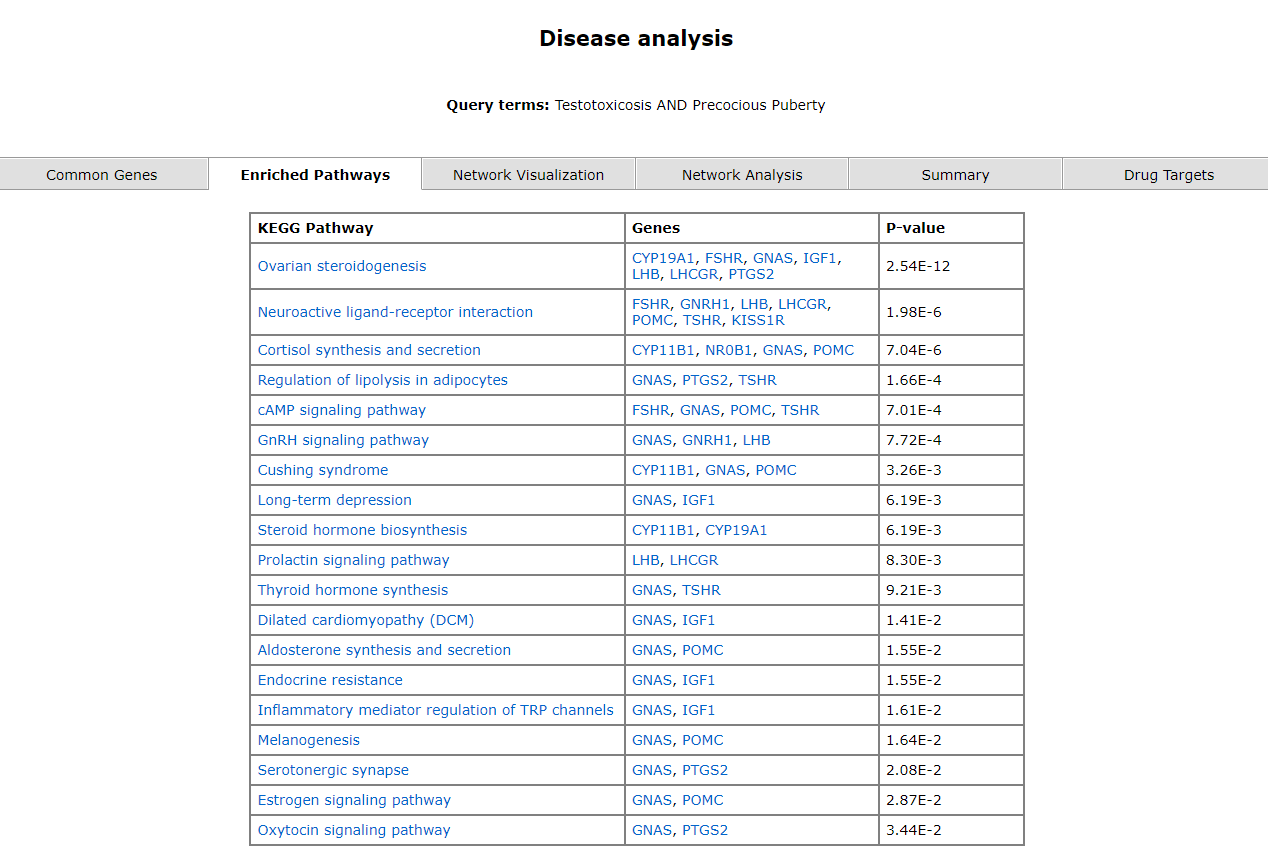
The disease analysis tab take the user to the pharmacological prediction tool. It has six tabs, each of which has the following data; common genes, enriched pathways, network visualization, network analysis, summary and drug targets.
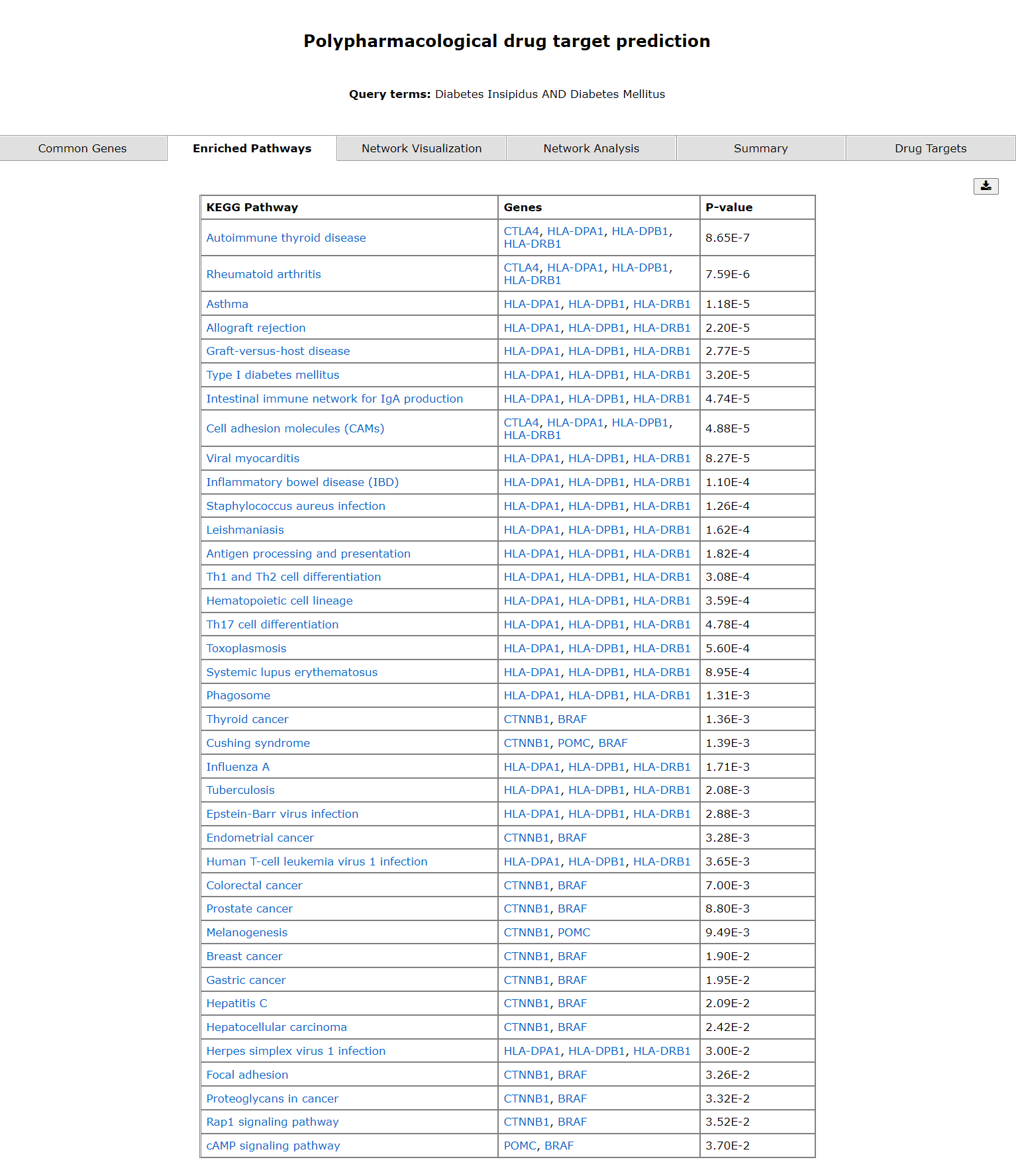
The common genes are mentioned in the first tab. The second tab displays a table of enriched pathways.
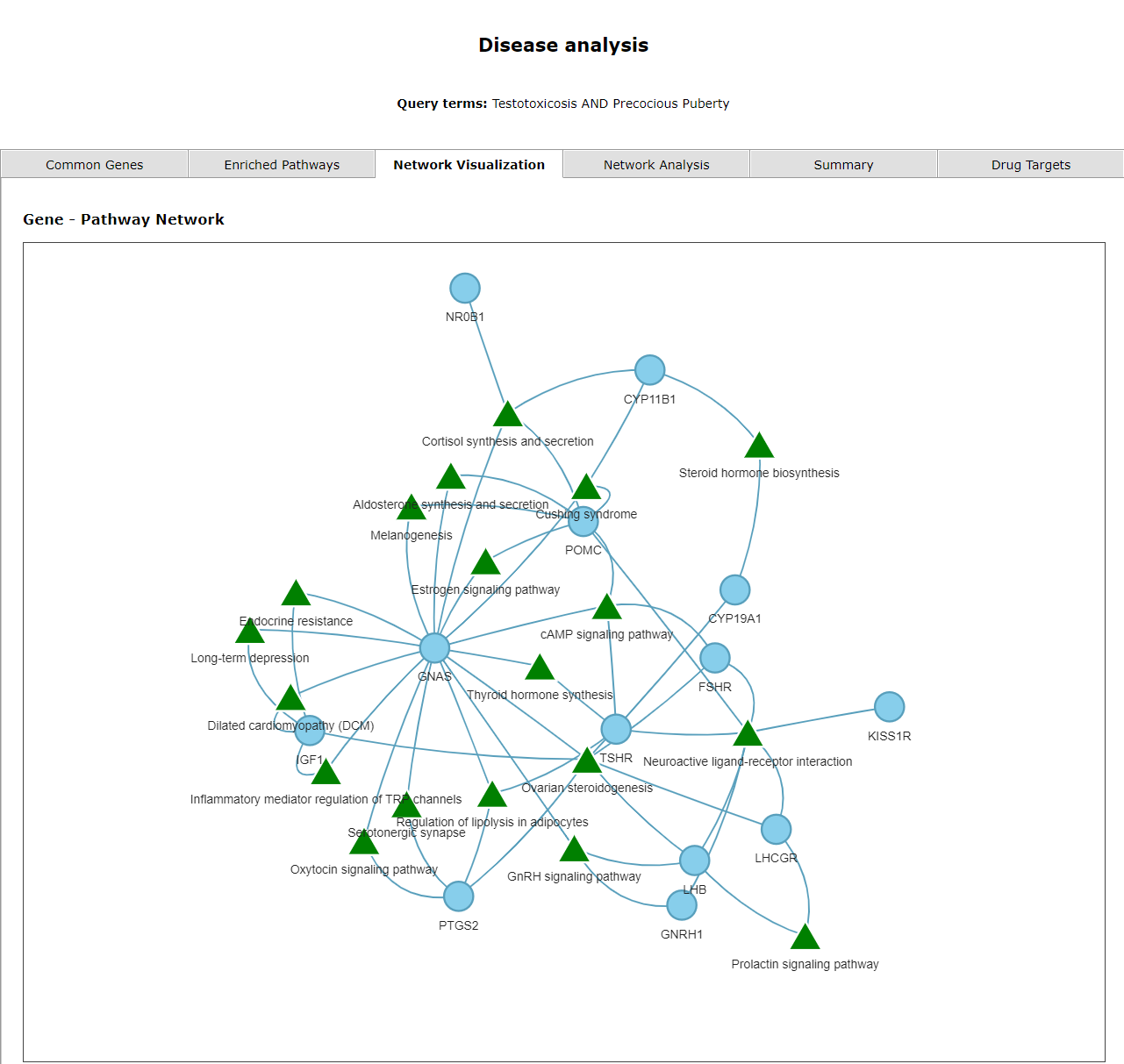
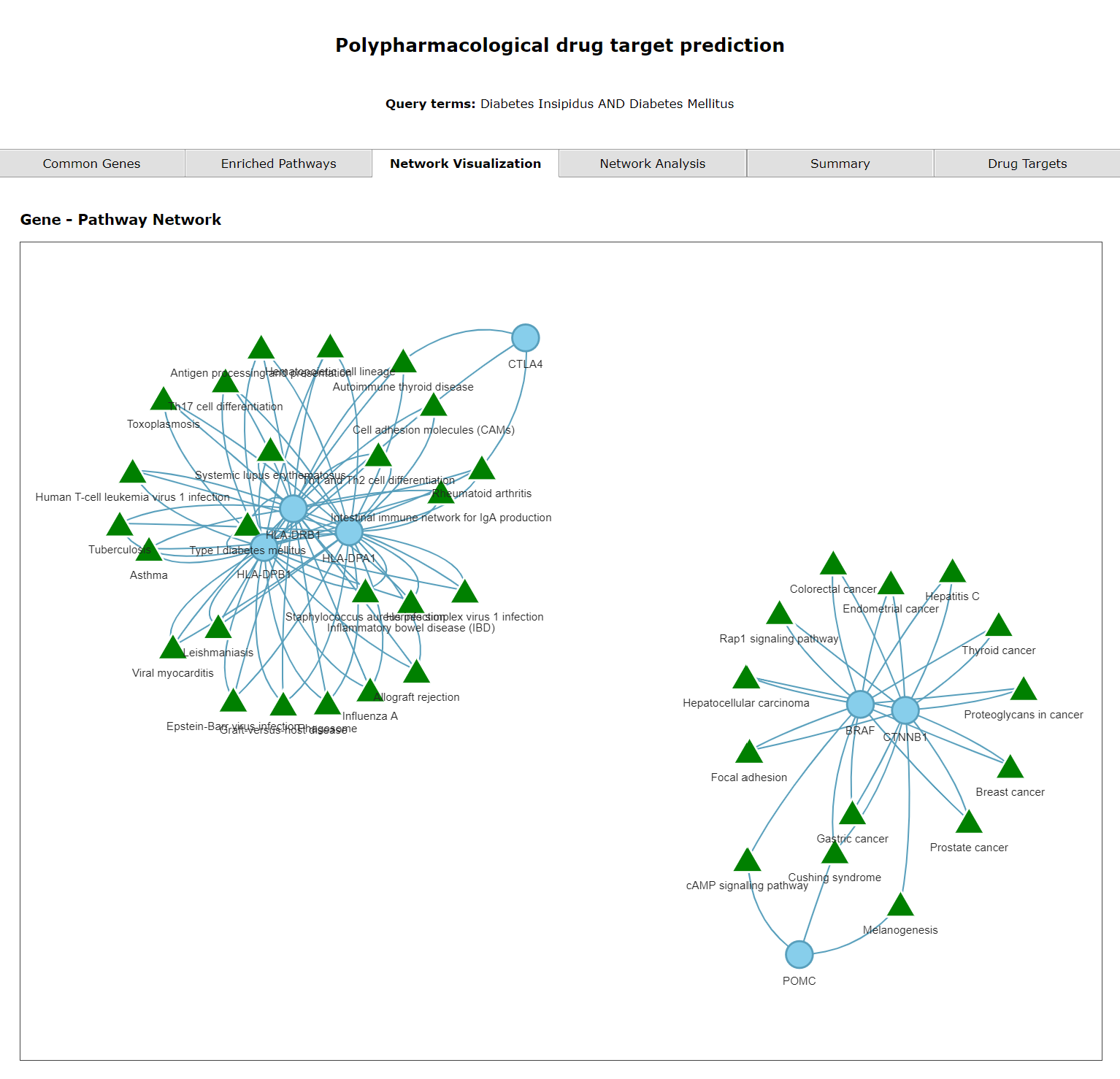
The third tab shows a network of enriched pathways and their genes corresponding to the selected diseases. The triangles represent pathways and circles represent genes.
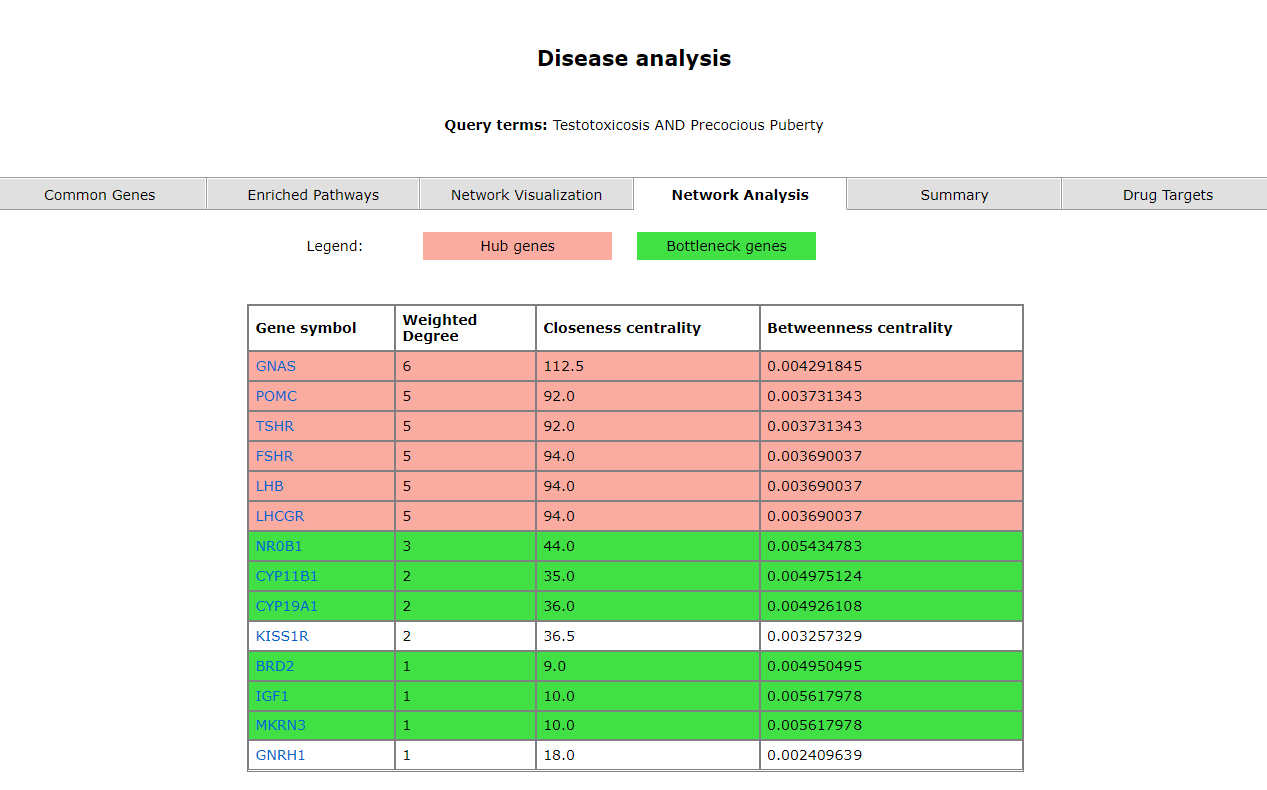
The network analysis tab represents the enriched genes (and bottleneck and hub genes) of the two diseases. They're displayed in the order of the three scores; weighted degree, closeness centrality and betweeness centrality.
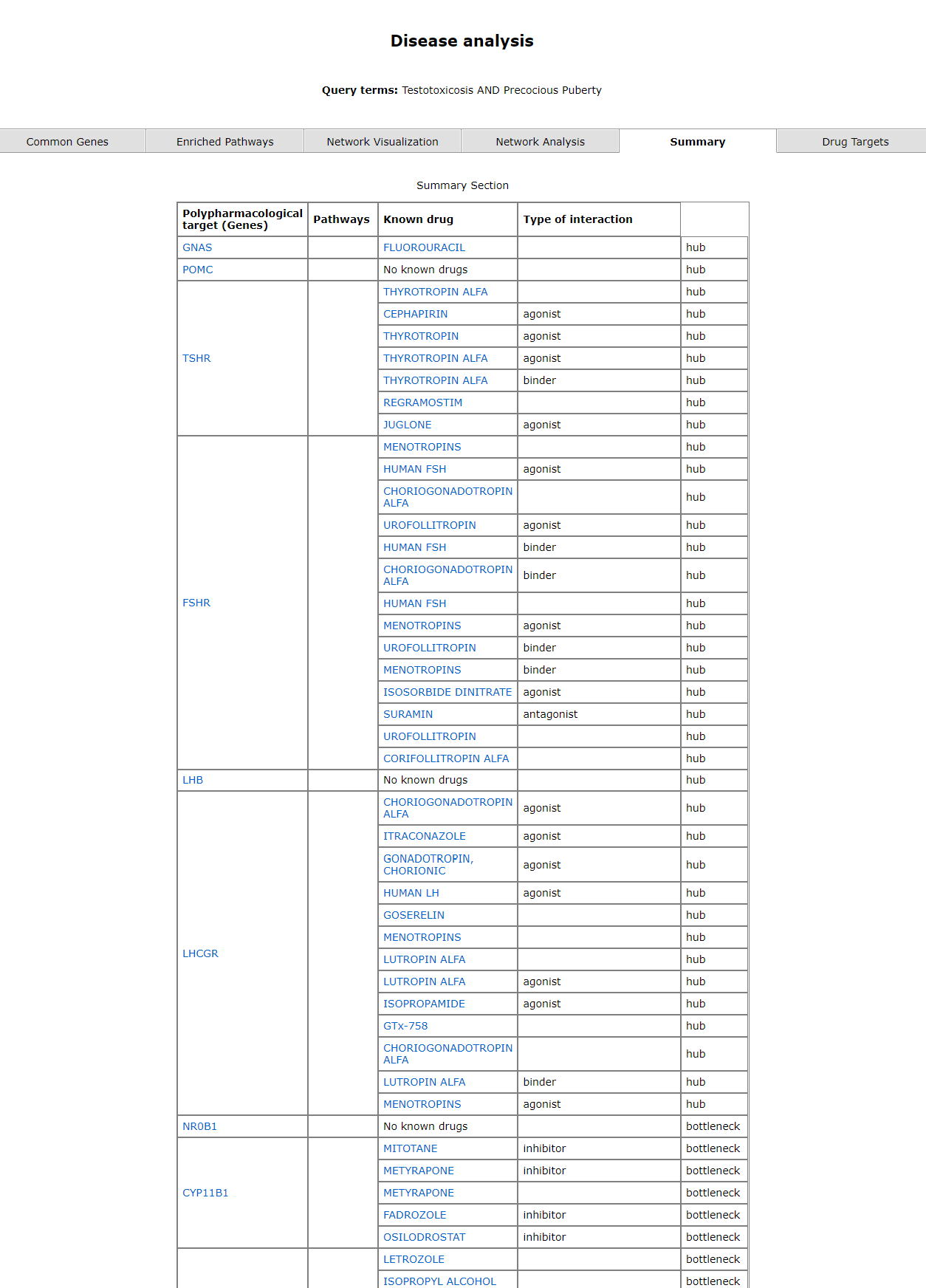
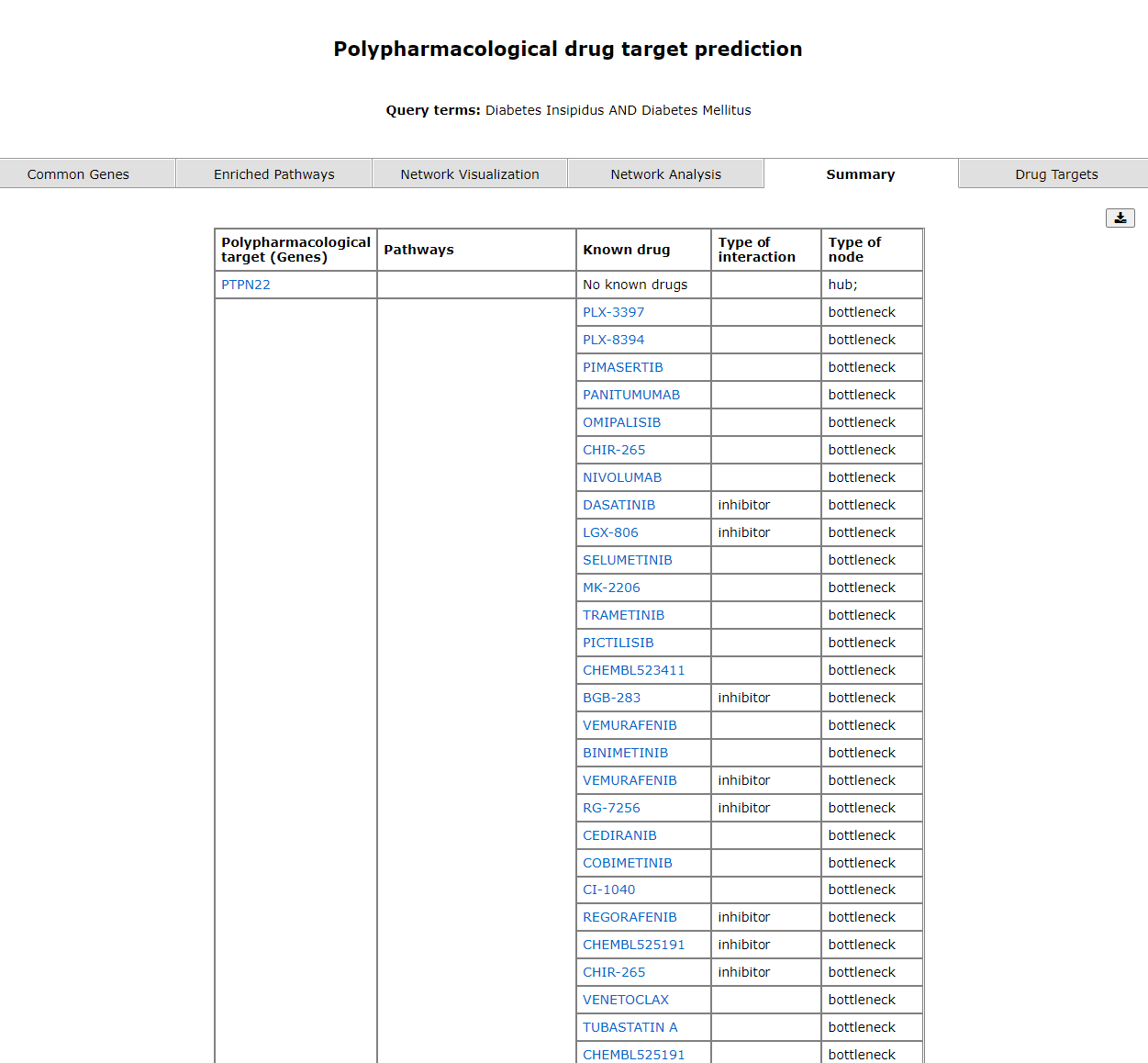
The summary tab gives a gist of the pharmacological target gene, known drug and its action (taken from https://www.dgidb.org/). The prediction of the gene being a bottleneck or hub is also mentioned.
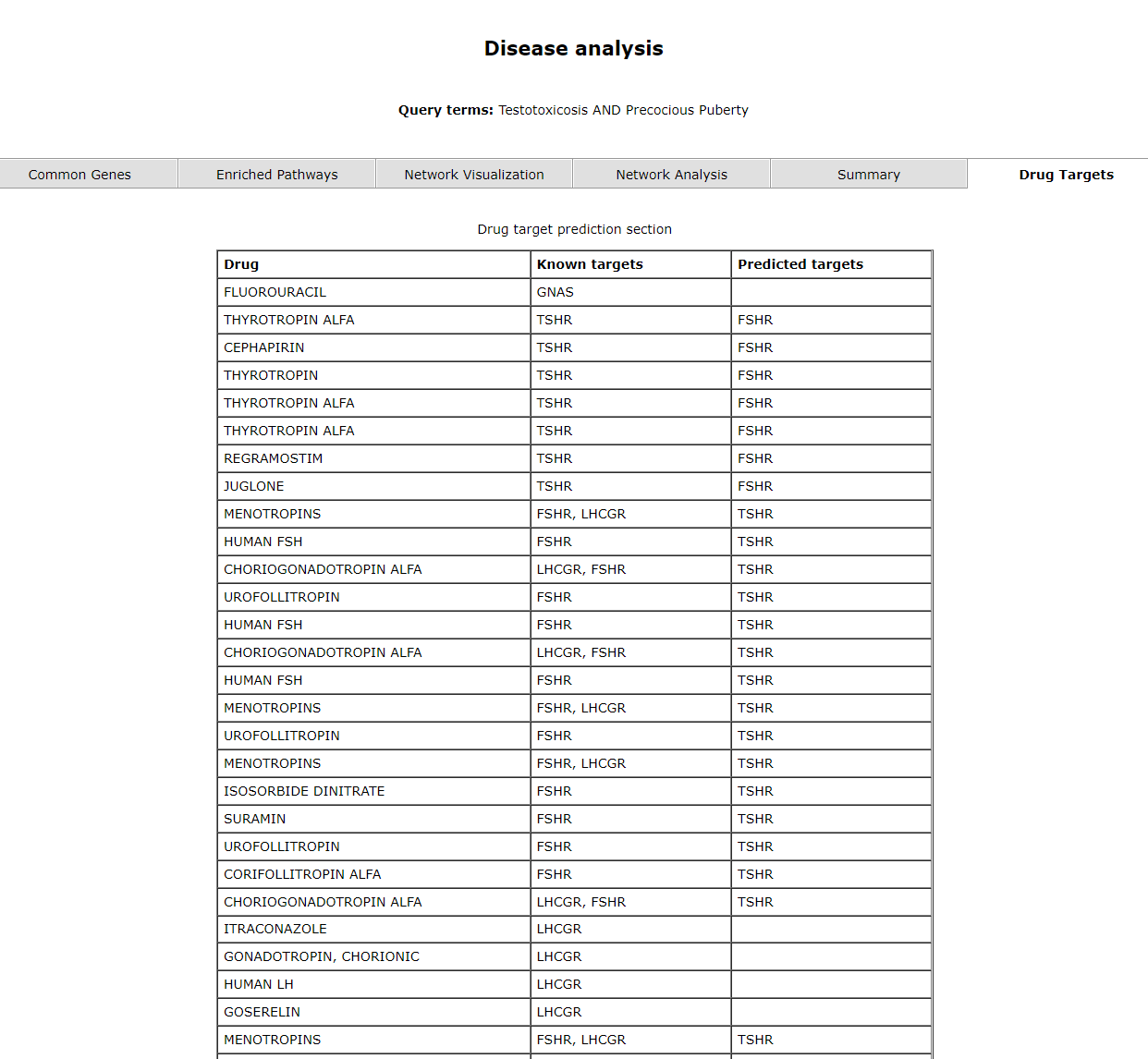
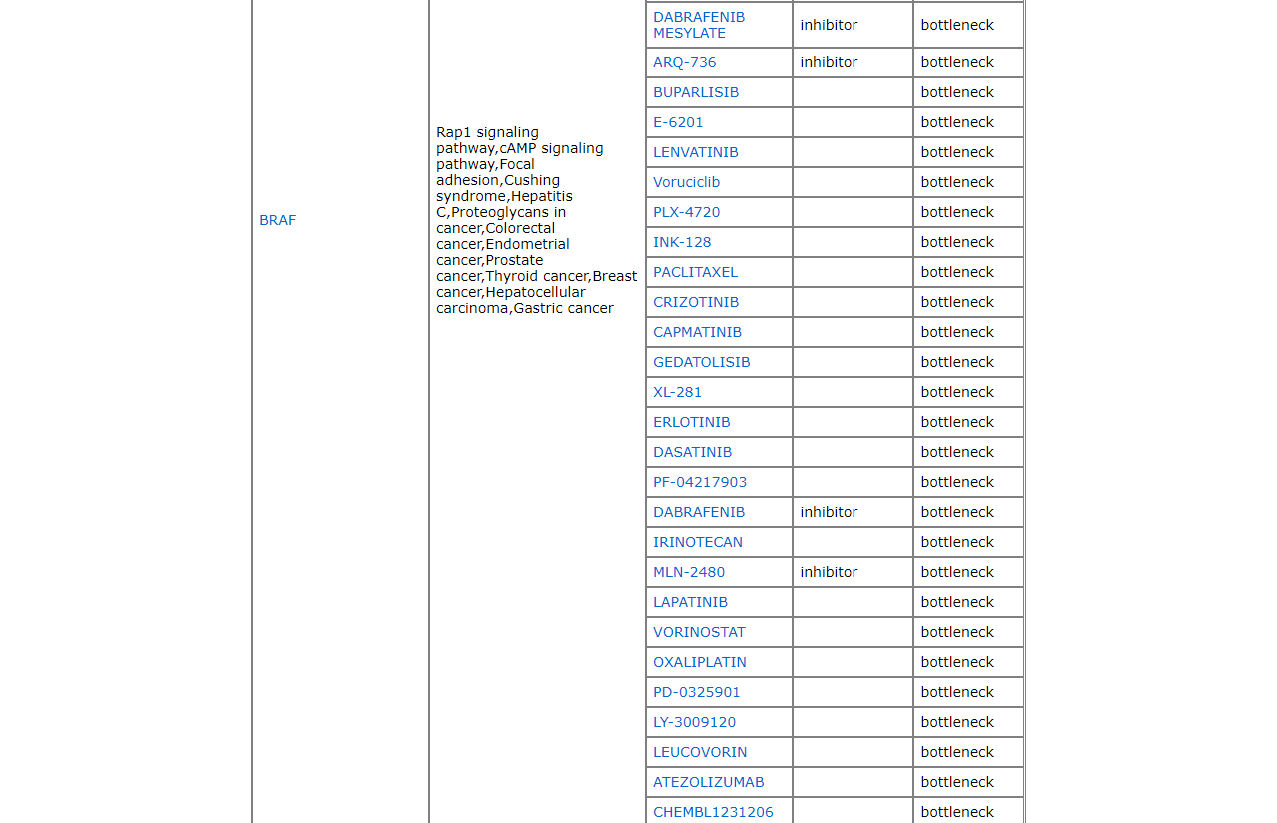
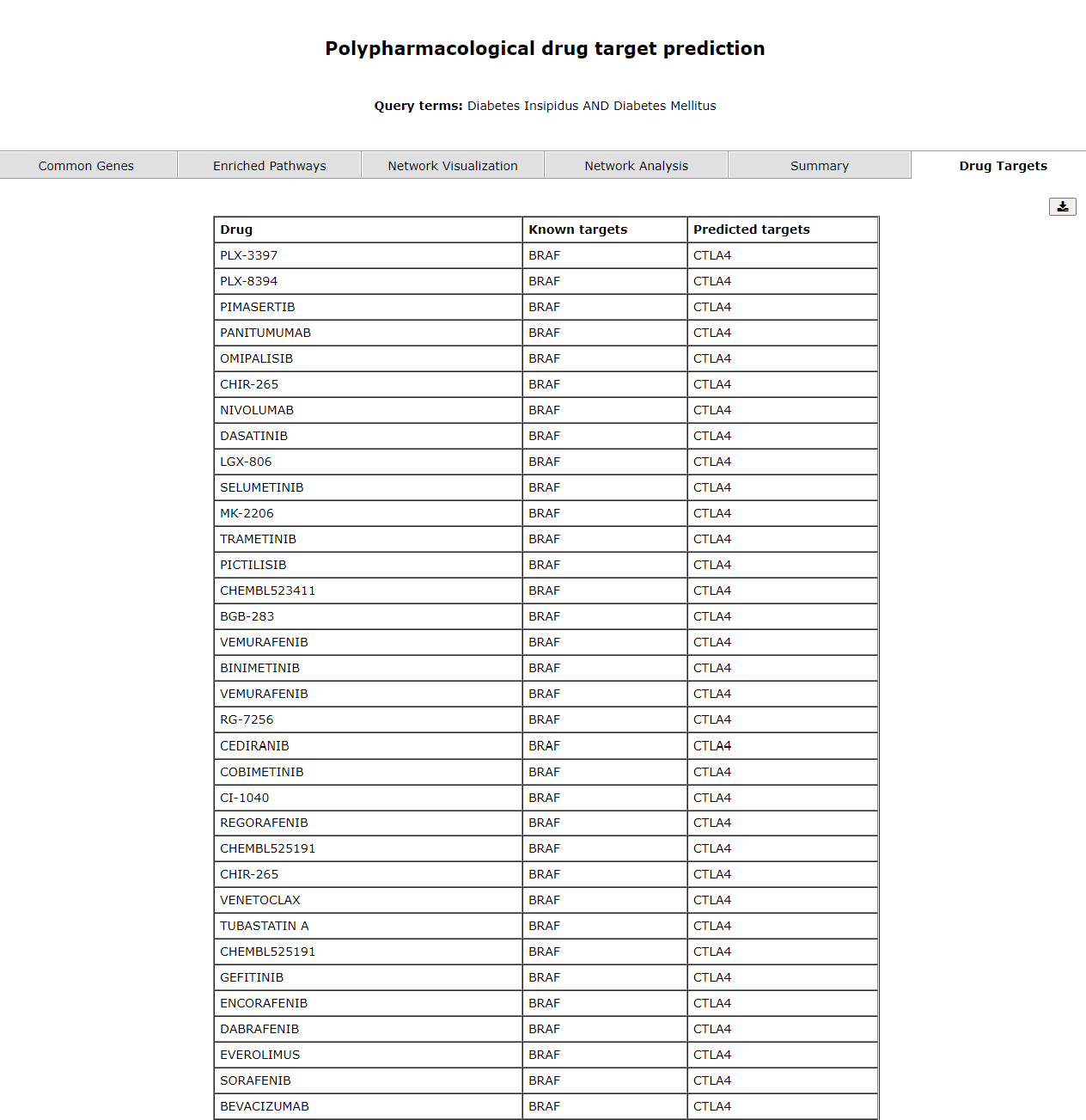
The last tab, drug targets provides user with a list of drugs, known targets and predicted targets. 4.2 Comorbidity Prediction
This tool can be used to predict comorbidity based on shared genes and their ontologies, uniqueness of shared genes, and network-based separation between them. For more details of the methodology refer to https://bmcmedgenomics.biomedcentral.com/articles/10.1186/s12920-017-0265-2. The home page of the tool has a list of diseases in the left column. User can select any number of disease names using the search with autofill option and use >> symbol to move the name in “selected diseases” column. Multiple diseases can be selected using "ctrl" key of the keyboard. An option to deselect diseases by using << symbol is also provided. Users can select from either of the two datasets, “Curated” and “Curated + Text mining”. 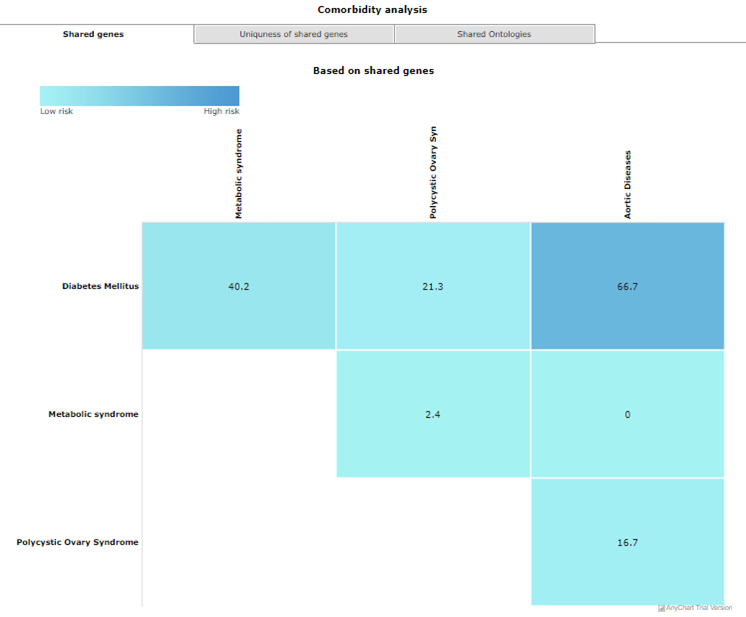
Results are displayed as heat maps. Darker the shade of blue higher is the comorbidity of the disease pair. Results can be downloaded using right click option. 4.3 Polypharmacological target prediction
Using this tool, potential polypharmacological drug targets and their known drugs from https://www.dgidb.org/ can be predicted.For more details of the methodology refer to (paper link). The homepage of the tool has a list of diseases. User can select any number of disease names using the search with autofill option and use >> symbol to move the name in “selected diseases” column. Multiple diseases can be selected using "ctrl" key of the keyboard. An option to deselect diseases by using << symbol is also provided. Users can select from either of the two datasets, “Curated” and “Curated + Text mining”. 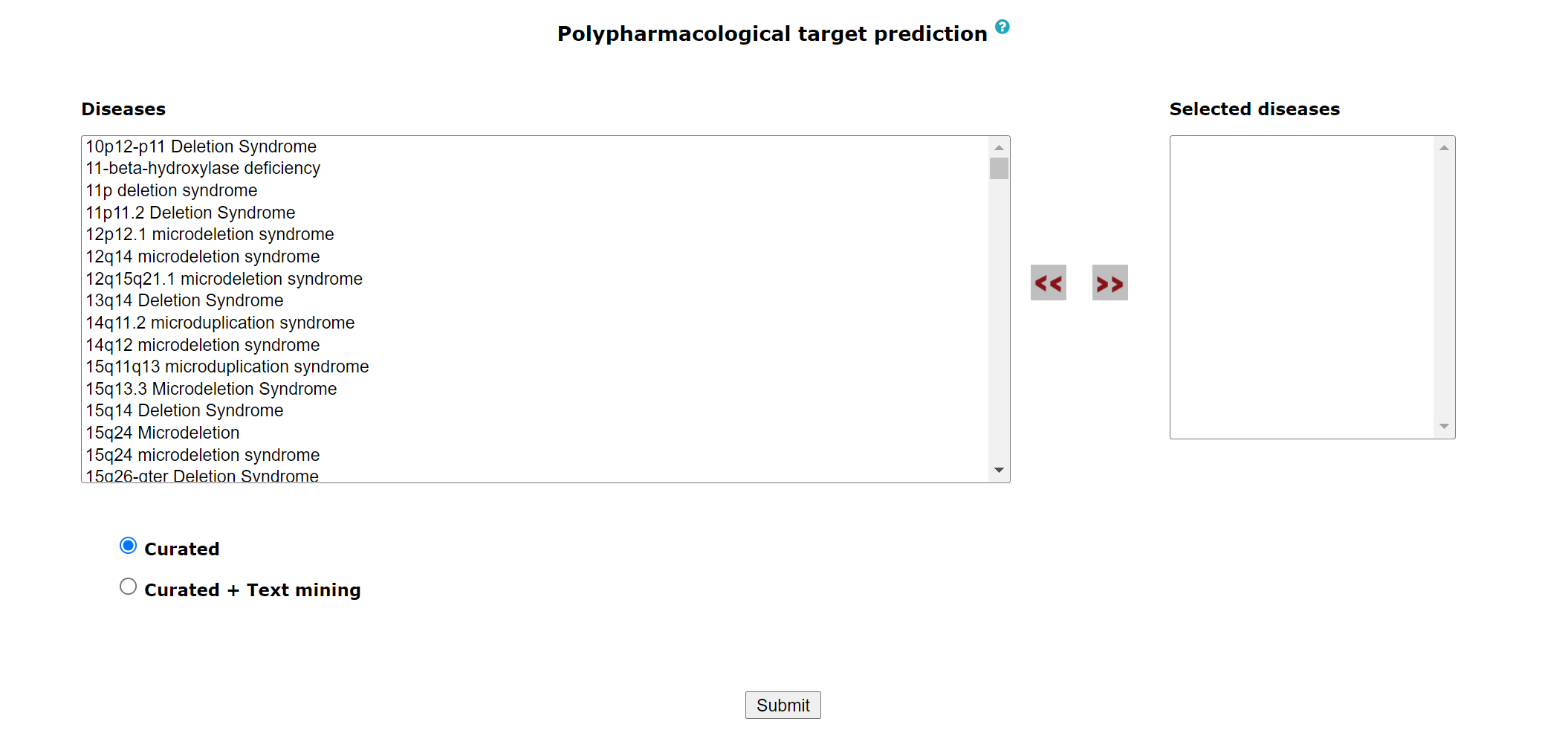
The submit button takes the user to the polypharmacological drug target prediction page. 
The output of the tool is as is mentioned in the disease analysis part of Section 4.1. The first tab shows common genes.
The second tab shows enriched pathways. The third tab shows gene pathway networks of the diseases.
The fourth tab displays the enriched hub and bottleneck genes along with their scores. 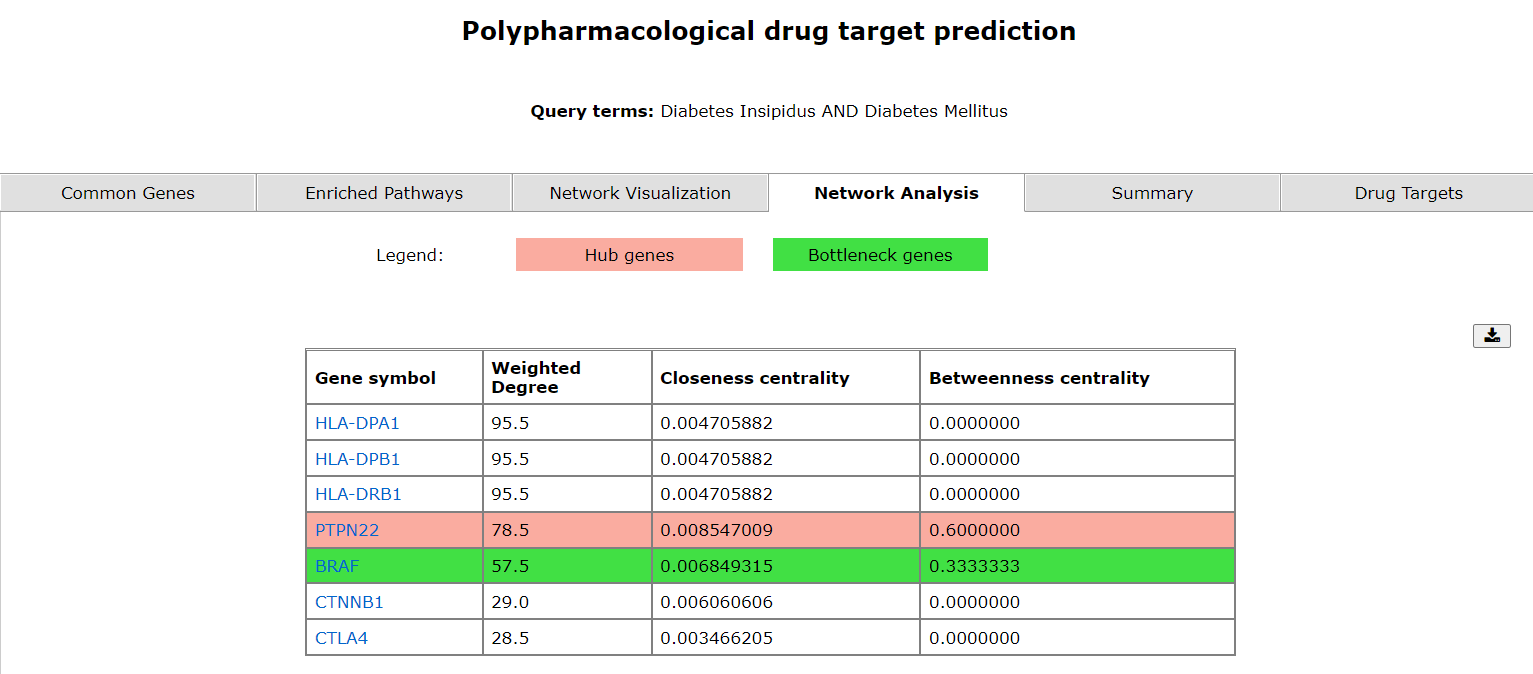
The fifth tab shows a summary of predicted drug targets.
The drug target page shows the predicted drug targets.
4.4 Venn Analysis
This tool can be used to illustrate the unique and/or common genes, pathways, ontologies, and Pfam domains for 2 or more (up to 6) diseases. A custom Venn for users to input their own data is also incorporated in the database. Home page for Venn looks like this
On selecting one type of Venn, user will be redirected to the main page of that Venn. Disease names can be entered in the The image below is how the example page of venn looks like. Top left of the page shows the venn diagram, with number of uniquely common elements. The numbers are hyperlinked to the box below; list of elements will pop up there. List can be copied from there to any editor. To search for elements in Venn, use the search in Venn box on the right side of Venn diagram. To download elements of Venn, the options
5. Links
The links tab contains links of various databases where the data is sourced from, the libraries used for algorithms or visualizations, and options for downloading data from the database. 6. Contact us
Contact information of the host institution is provided in the “Contact us” tab. 7. Search icon
Search icon (small magnifying glass with a search bar 8. Save query
Clipboard icon ( |
| © 2021, Biomedical Informatics Centre, NIRRH |
ICMR-National Institute for Research in Reproductive Health, Jehangir Merwanji Street, Parel,
Mumbai-400012
Tel: +91-22-24192104, Fax No: +91-22-24139412

 button at the end of the web page.Pathways can be selected using the checkbox option against each pathway name. The selection can be reset using the
button at the end of the web page.Pathways can be selected using the checkbox option against each pathway name. The selection can be reset using the  back button takes the user to the initial page of enrichment analysis.
back button takes the user to the initial page of enrichment analysis. button. It will result in a network of selected genes.
button. It will result in a network of selected genes.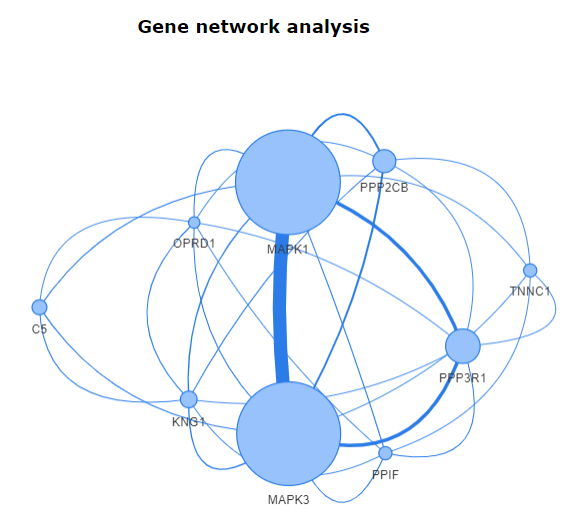
 button at the end of the web page.
button at the end of the web page.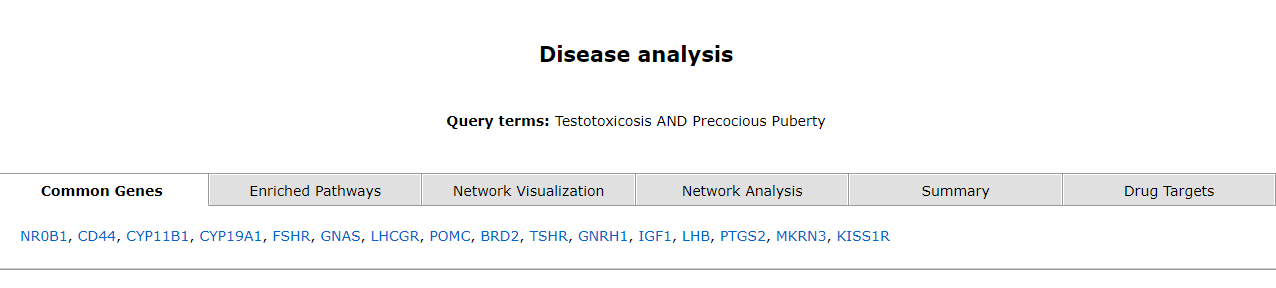
 button can be used to go back to the input page.
button can be used to go back to the input page.